# Add two replicated observations per plant # Model with 2 plants at with the same row/col position M2 <- asreml(y ~ 1,rcov=~iexp(row,col),random=~plant,na.method.X="include",data=data1,trace=F) # Should work even when the grid is not completeĭata1$y <- NA # Matern with some missing plants in the grid # Matérn model equivalent to isotropic AR1 (equivalent to m3 with lambda = "1F") M2 <- asreml(y ~ 1,rcov=~iexp(row,col),random=~plant,data=data,trace=F) # Isotropic model with the exponential function M1 <- asreml(y ~ 1,rcov=~exp(rowfac):exp(colfac),random=~plant,data=data,trace=F) M0 <- asreml(y ~ 1,rcov=~ar1(rowfac):ar1(colfac),random=~plant,data=data,trace=F) # Dataset should be with columns nested within rows

# Compute euclidian distance between each data point (ie each row)ĭist <- as.matrix(dist(cbind(data$row,data$col))) # Add spatially correlated and spatially uncorrelated errorĪutocorrerror <- <- rnorm(nrow(data),0,siguncorres)ĭata$y <- intercept+autocorrerror+uncorrerror Model <- RMexp(scale=-1/log(rho), var=sigcorres^2) # Initialize AR1 simulation model using the spatial correlation parameter rho and the variance for spatially correlated effects # Create dataset with 20 row and 20 columns #install.packages(packageurl, repos=NULL, type="source")
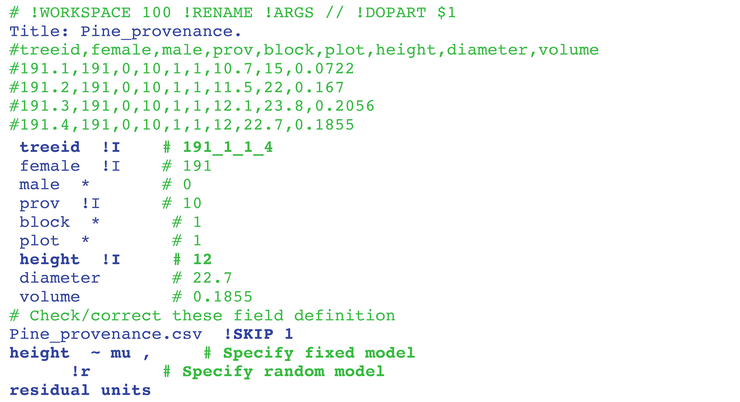
ASREML FREE VERSION INSTALL
# To install the version of the package 'proxy' compatible with R.3.2.2 # for a more deteiled presentation, see: # Goal of this document: Compare AR1, exponential and Matern models (with between spatially correlated error variance and spatially uncorrelated (nugget) error variance using ASReml-R/spaMM
